Selected websites dedicated to Covid-19
The web site of the Federation of European Societies of Microbiology has two dedicated web pages: on the novel coronavirus and the expert update.
The news web page of the Imperial College, London, though not explicitly dedicated to the current pandemics, has several relevant news on it.
The London School of Hygiene and Tropical Medicine has a web page on Covid-19.
The web site of the European Centre for Disease Control hosts a page that reports the daily statistics of the pandemics.
The World Health Organization has a page on Covid-19 and pubblishes the situation reports with almost daily frequence.
The Center for Disease Control, Atlanta, GA, USA.
The RCSB Protein Data Bank website has a webpage on structural biology research on Coronavirus.
The UniProt website has a webpage that reports the annotated sequences of proteins from Covid-19.
All major medical journals have pages dedicated to Covid19:
British Medical Journal
JAMA
NEJM
The Lancet
image courtesy of Pixabay
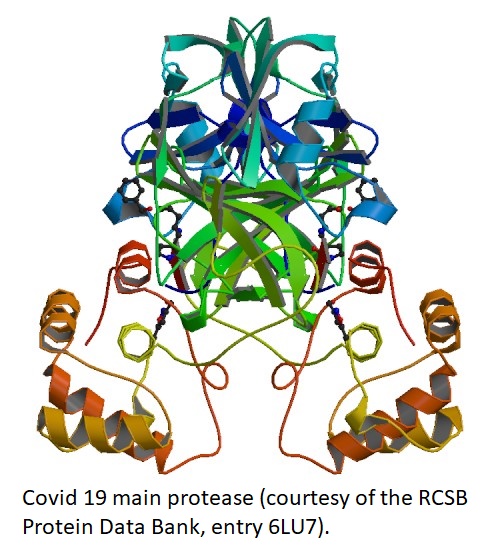
Zhang L, Lin D, Sun X, Curth U, Drosten C, Sauerhering L, Becker S, Rox K, Hilgenfeld R. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science. 2020 Mar 20. pii: eabb3405. doi: 10.1126/science.abb3405. Structure of and binding of specific inhibitors to the main protease from Covid-19. This protease is a symmetric homodimer, whose catalytic residues are His41 and Cys145. The two catalytic sites are on oposite sides of the dimer, far from one another and from the monomer-monomer interface.
Wang L, Hu W, Fan C. Structural and biochemical characterization of SADS-CoV papain-like protease 2. Protein Sci. 2020 Mar 26. doi: 10.1002/pro.3857. Processing of the polyproteins pp1a and pp1ab produced by coronaviruses requires at least two or three specific viral proteases. This paper reports the structure and properties of the papain-like protease from a swine coronavirus related to Covid-19.
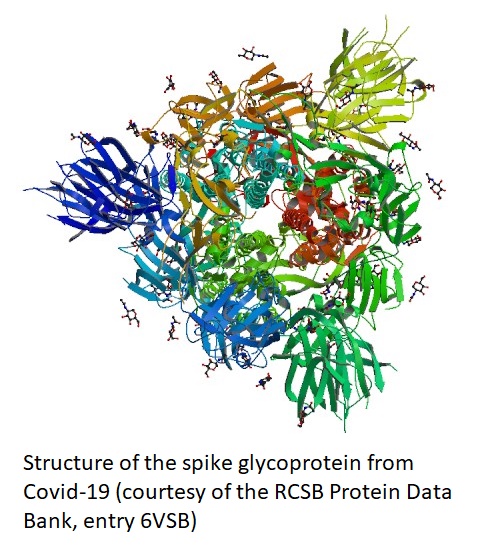
Lan J, Ge J, Yu J, Shan S, Zhou H, Fan S, Zhang Q, Shi X, Wang Q, Zhang L, Wang X. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020 Mar 30. doi: 0.1038/s41586-020-2180-5.
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020 Mar 13;367(6483):1260-1263. doi: 10.1126/science.abb2507.
Mallapaty S. Why does the coronavirus spread so easily between people? Nature. 2020 Mar;579(7798):183. doi: 10.1038/d41586-020-00660-x. A summary of the biochemical mechanisms of Covid-19 binding to ACE2 on the host cell membrane and its proteolytic activation by furin, which explains the broad organ specificity of the virus.
Hussain M, Jabeen N, Raza F, Shabbir S, Baig AA, Amanullah A, Aziz B. Structural Variations in Human ACE2 may Influence its Binding with SARS-CoV-2 Spike Protein. J Med Virol. 2020 Apr 6. doi: 10.1002/jmv.25832. Genetic variants of ACE2, the human protein that acts as the receptor for Covid-19, may confer resistance to the infection. This has important implications for the predictions of the possible attack rate of the pandemics and the public policy decisions on its containment.
Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, Guo L, Guo R, Chen T, Hu J, Xiang Z, Mu Z, Chen X, Chen J, Hu K, Jin Q, Wang J, Qian Z Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun. 2020 Mar 27;11(1):1620. doi: 10.1038/s41467-020-15562-9.
Shang J, Ye G, Shi K, Wan Y, Luo C, Aihara H, Geng Q, Auerbach A, Li F. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science. 2020 Mar 27;367(6485):1444-1448. doi: 10.1126/science.abb2762.
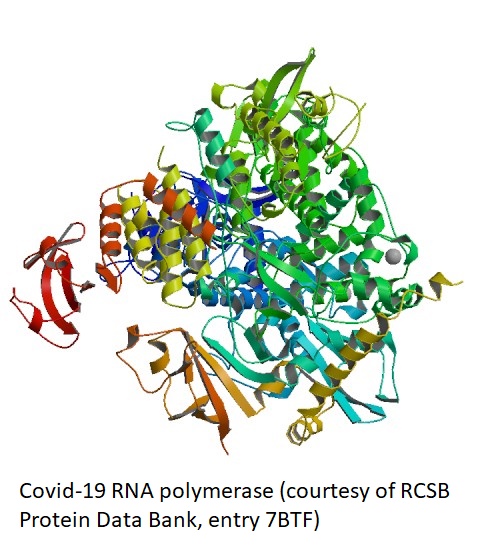
Gordon CJ, Tchesnokov EP, Feng JY, Porter DP, Gotte M. The antiviral compound remdesivir potently inhibits RNA-dependent RNA polymerase from Middle East respiratry syndrome coronavirus. J Biol Chem. 2020 Feb 24. pii: jbc.AC120.013056. doi: 10.1074/jbc.AC120.013056. Remdesvir is synthetic nucleotide analog and a selective inhibitor of viral RNA-dependent RNA polymerases. This paper demonstrates thath it inhibitits the replication of the MERS Covid, a virus species related to Covid-19.
Yan Gao, Liming Yan, Yucen Huang, Fengjiang Liu, Yao Zhao, Lin Cao, Tao Wang, Qianqian Sun, Zhenhua Ming, Lianqi Zhang, Ji Ge, Litao Zheng, Ying Zhang, Haofeng Wang, Yan Zhu, Chen Zhu, Tianyu Hu, Tian Hua, Bing Zhang, Xiuna Yang, Jun Li, Haitao Yang, Zhijie Liu, Wenqing Xu, Luke W. Guddat, Quan Wang, Zhiyong Lou, Zihe Rao Structure of RNA-dependent RNA polymerase from 2019-nCoV, a major antiviral drug target bioRXiv, doi: https://doi.org/10.1101/2020.03.16.993386. This paper reports the cryo-EM structure of 2019-nCoV full-length RNA-dependent RNA polymerase (protein nsp12) in complex with cofactors nsp7 and nsp8 at a resolution of 2.9 A.
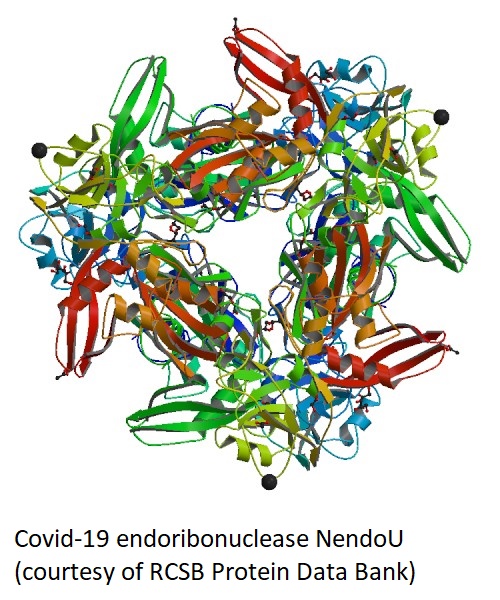
Youngchang Kim, Robert Jedrzejczak, Natalia I. Maltseva, Michael Endres, Adam Godzik, Karolina Michalska, Andrzej Joachimiak Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2 bioRXiv doi: https://doi.org/10.1101/2020.03.02.968388
Kumar S, Maurya VK, Prasad AK, Bhatt MLB, Saxena SK. Structural, glycosylation and antigenic variation between 2019 novel coronavirus (2019-nCoV) and SARS coronavirus (SARS-CoV). Virusdisease. 2020 Mar;31(1):13-21. doi: 10.1007/s13337-020-00571-5.
Tai W, He L, Zhang X, Pu J, Voronin D, Jiang S, Zhou Y, Du L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol. 2020 Mar 19. doi: 10.1038/s41423-020-0400-4.
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020 Feb 22;395(10224):565-574. doi: 10.1016/S0140-6736(20)30251-8. Epub 2020 Jan 30.
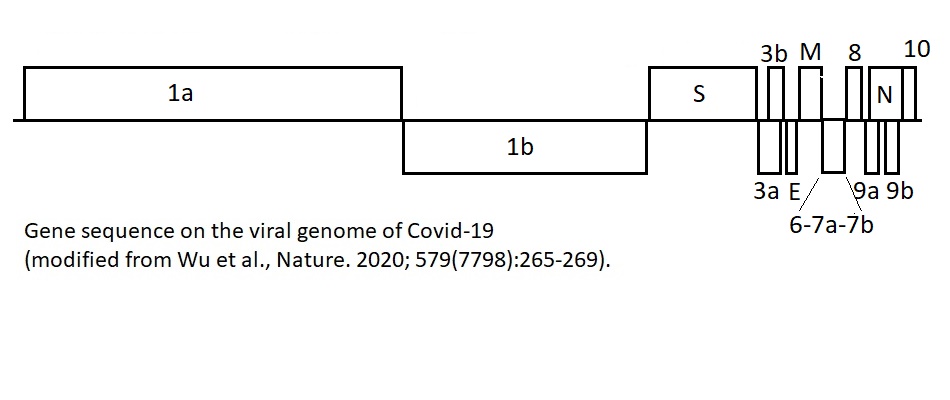
Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, Hu Y, Tao ZW, Tian JH, Pei YY, Yuan ML, Zhang YL, Dai FH, Liu Y, Wang QM, Zheng JJ, Xu L, Holmes EC, Zhang YZ. A new coronavirus associated with human respiratory disease in China. Nature. 2020 Mar;579(7798):265-269. doi: 10.1038/s41586-020-2008-3.
Mousavizadeh L, Ghasemi S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J Microbiol Immunol Infect. 2020 Mar 31. pii: S1684-1182(20)30082-7. doi: 10.1016/j.jmii.2020.03.022.
Ferguson et al.Impact of non-pharmaceutical interventions (NPIs) to reduce COVID19 mortality and healthcare demand. Document of March 16, 2020 by the Imperial College COVID-19 Response Team.
Nicholas G. Davies, Adam J. Kucharski, Rosalind M. Eggo, Amy Gimma, CMMID COVID-19 working group, W. John Edmunds The effect of non-pharmaceutical interventions on COVID-19 cases, deaths and demand for hospital services in the UK: a modelling study. Report of the London School of Hygiene & Tropical Medicine
Ioannidis JPA. Coronavirus disease 2019: The harms of exaggerated information and non-evidence-based measures. Eur J Clin Invest. 2020 Apr;50(4):e13222. doi: 10.1111/eci.13222
Verity R. et al. Estimates of the severity of coronavirus disease 2019: a model-based analysis. Lancet Infect Dis. 2020. pii: S1473-3099(20)30243-7. doi: 10.1016/S1473-3099(20)30243-7.
Li Yang Hsu, Po Ying Chia, Jeremy FY Lim The Novel Coronavirus (SARS-CoV-2) Epidemic. Annals, Academy of Medicine, Singapore, 2020.
Covid-19: the CIDRAP viewpoint; Moore, Lipsitch, Barry, Osterholm Part 1: The Future of the COVID-19 Pandemic: Lessons Learned from Pandemic Influenza.
Rice K, Wynne B., Martin V, and Ackland GJ: Effect of school closures on mortality from coronavirus disease 2019: old and new predictions (BMJ 2020; 371). A reevaluation on the estimates of mortality and the effect of containment measures. Take home message: whatever measures one takes, the mortality will depend on the most affected age groups, and the epidemic ends when herd immunity is reached. Closing schools may cause increased overall mortality because it reduces the burden on age groups affected by mildes disease.